Lab 4
Decision Trees, Bagging, Random Forests, and Boosting
Decision Trees
Fitting Classification Trees
- The
treelibrary in R is used to construct regression trees
- Fit a regression tree using the
Bostondataset
- Split data into a training set
- Train the tree model on the training data
library(MASS)
set.seed(1)
train <- sample(1:nrow(Boston), nrow(Boston) / 2)
tree.boston <- tree(medv ~ ., Boston, subset = train)
summary(tree.boston)
Regression tree:
tree(formula = medv ~ ., data = Boston, subset = train)
Variables actually used in tree construction:
[1] "rm" "lstat" "crim" "age"
Number of terminal nodes: 7
Residual mean deviance: 10.38 = 2555 / 246
Distribution of residuals:
Min. 1st Qu. Median Mean 3rd Qu. Max.
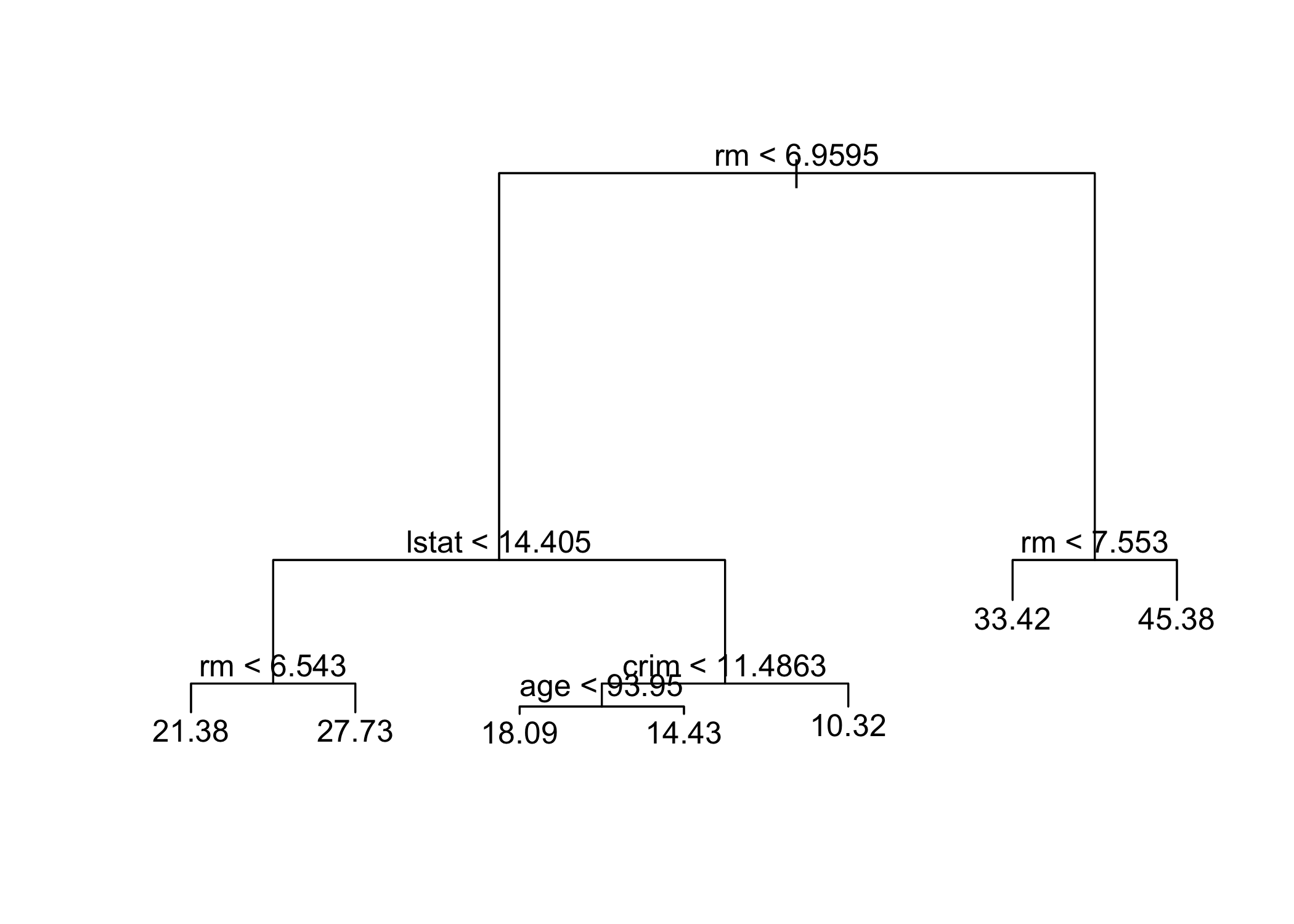
-10.1800 -1.7770 -0.1775 0.0000 1.9230 16.5800 summary()shows only 4 variables are used in the tree
- In regression trees, deviance = sum of squared errors (SSE)
lstat: % of lower socioeconomic status
rm: average number of roomsInterpretation:
- Higher
rm→ higher house prices
- Lower
lstat→ higher house prices
- Example:
rm ≥ 7.553→ predicted price ≈ $45,400
- Higher
Model flexibility:
- Larger tree can be grown using
tree.control(..., mindev = 0)
- Larger tree can be grown using
Next step:
- Use
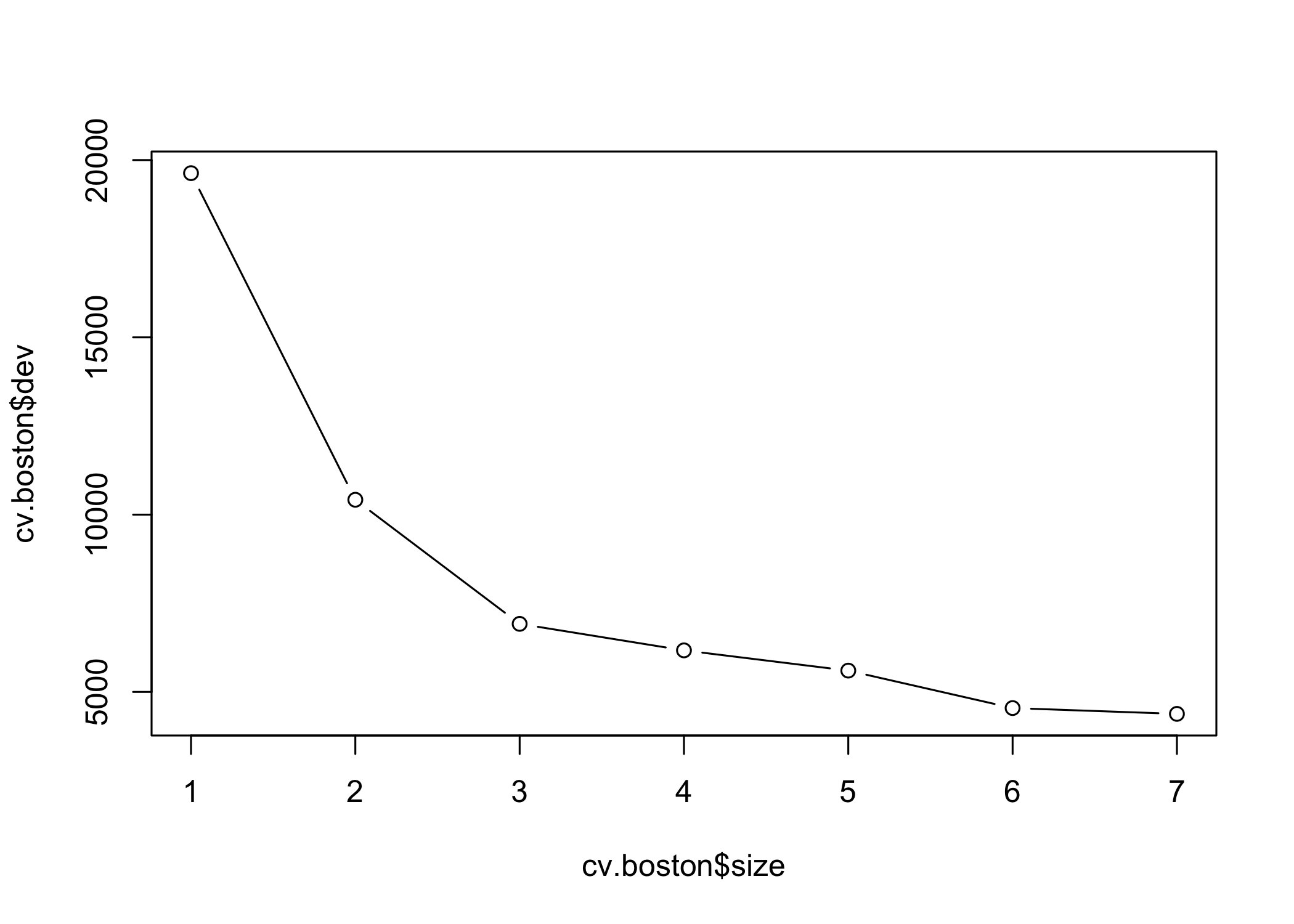
cv.tree()to evaluate pruning via cross-validation
- Use
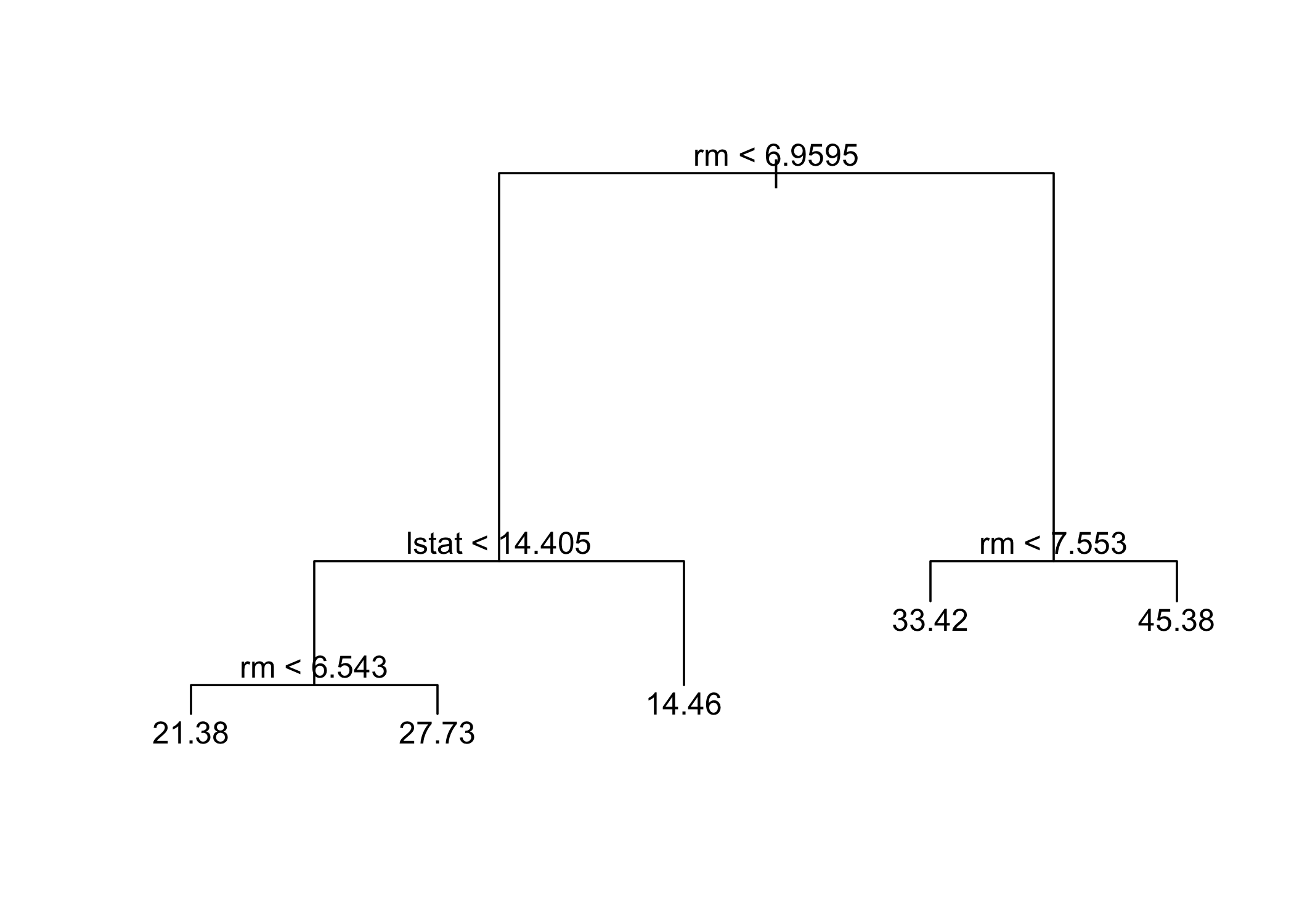
- Cross-validation selects the most complex tree in this case
- Tree can still be pruned manually if desired
- Based on cross-validation, the unpruned tree is selected
- Use the unpruned tree to make predictions on the test set
yhat <- predict(tree.boston, newdata = Boston[-train, ])
boston.test <- Boston[-train, "medv"]
plot(yhat, boston.test)
abline(0, 1)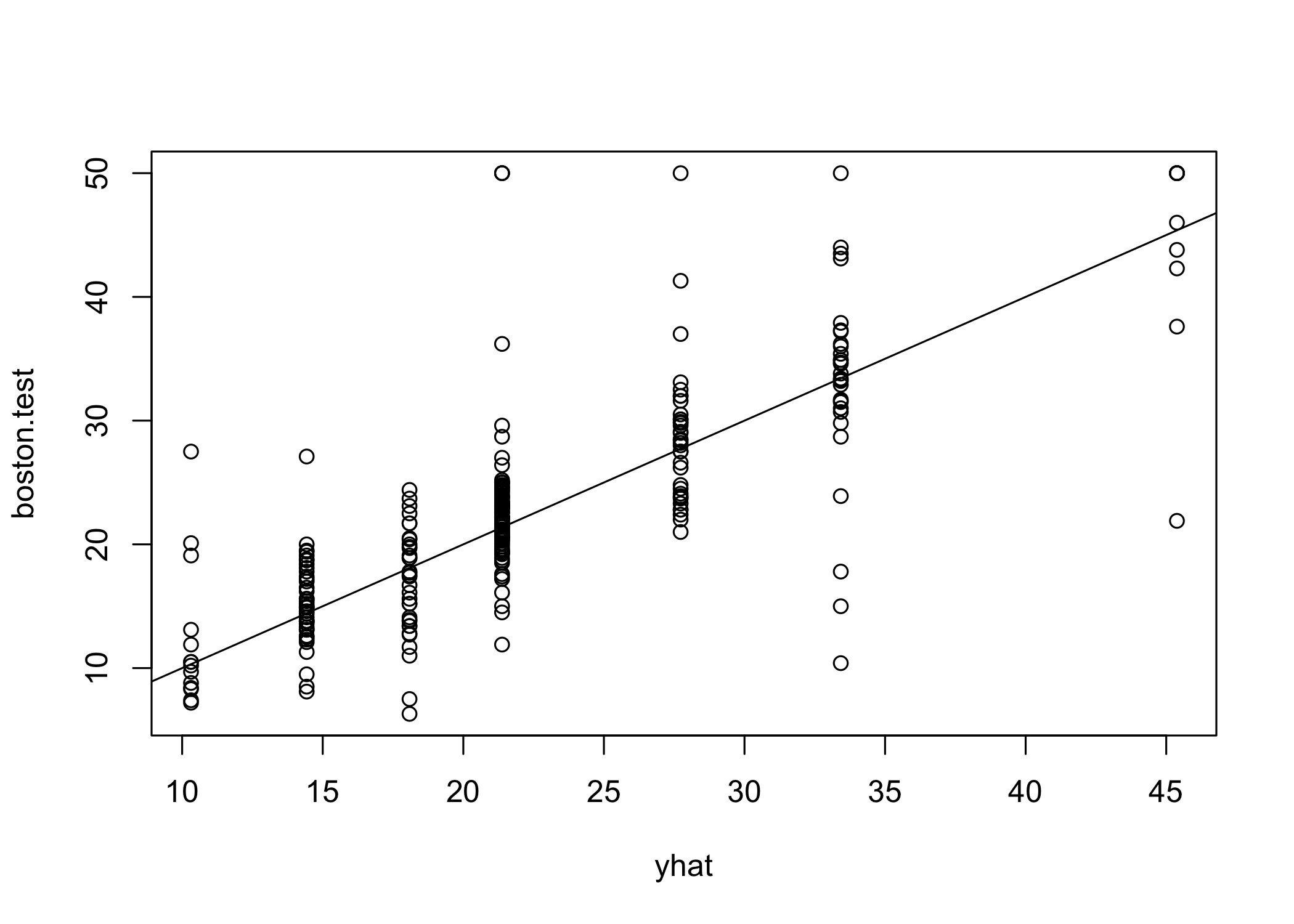
[1] 35.28688- Test set MSE ≈ 35.29
- RMSE ≈ 5.941
- Predictions are on average within ~$5,941 of true median house values
Bagging and Random Forests
- Apply bagging and random forests using the
randomForestpackage in R
- Bagging = special case of random forest with m = p (all predictors used at each split)
randomForest 4.7-1.1Type rfNews() to see new features/changes/bug fixes.set.seed(1)
bag.boston <- randomForest(medv ~ ., data = Boston,
subset = train, mtry = 12, importance = TRUE)
bag.boston
Call:
randomForest(formula = medv ~ ., data = Boston, mtry = 12, importance = TRUE, subset = train)
Type of random forest: regression
Number of trees: 500
No. of variables tried at each split: 12
Mean of squared residuals: 11.10176
% Var explained: 85.56mtry = 12→ all 12 predictors used at each split
- This corresponds to bagging
- Evaluate model performance on the test set
yhat.bag <- predict(bag.boston, newdata = Boston[-train, ])
plot(yhat.bag, boston.test)
abline(0, 1)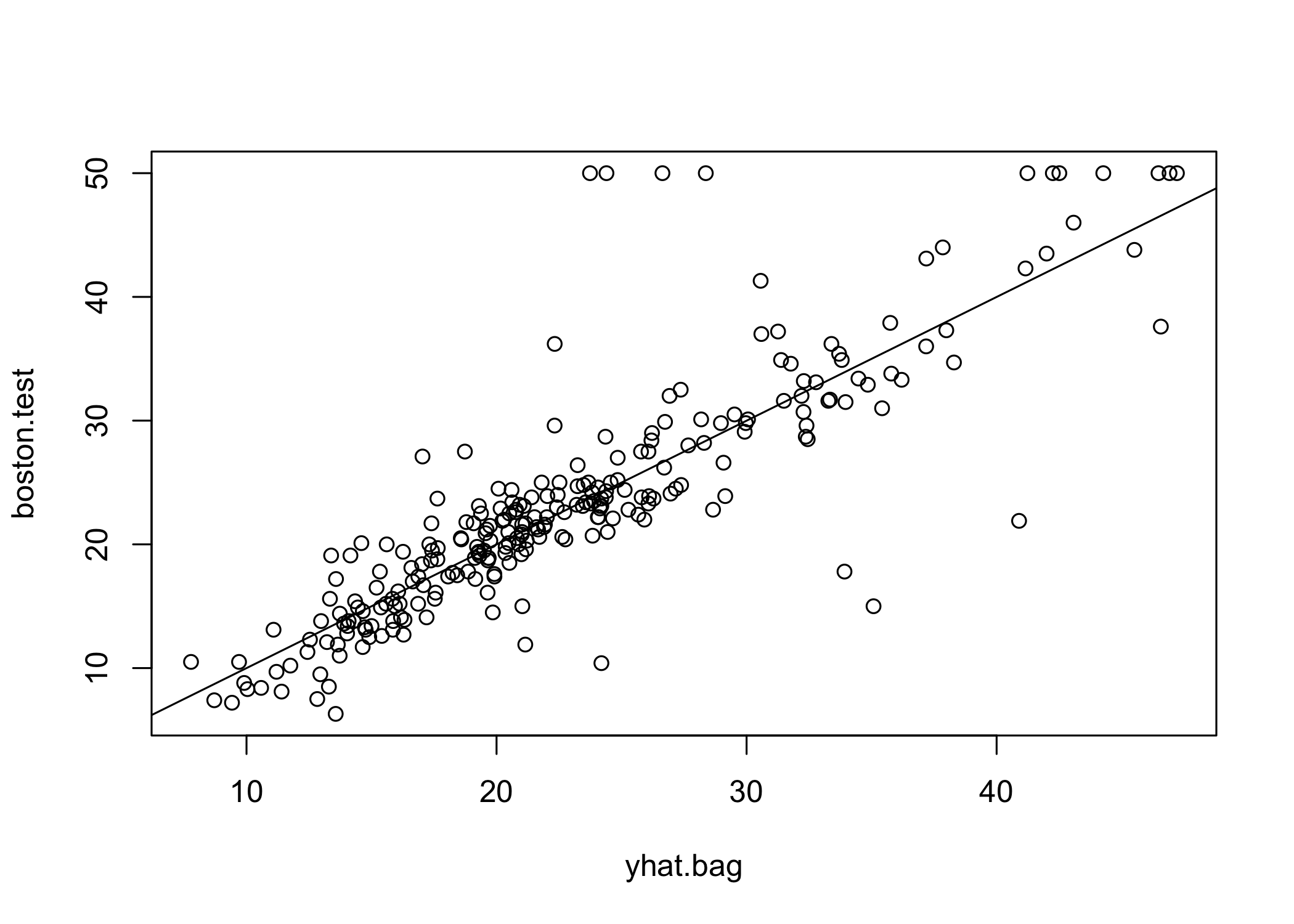
[1] 23.38773- Bagged model test MSE ≈ 23.42
- Significantly lower than single tree (~2/3 of its error)
- Number of trees can be controlled with the
ntreeparameter
bag.boston <- randomForest(medv ~ ., data = Boston,
subset = train, mtry = 12, ntree = 25)
yhat.bag <- predict(bag.boston, newdata = Boston[-train, ])
mean((yhat.bag - boston.test)^2)[1] 25.19144- Random forests are built like bagging but with smaller
mtry
- Default
mtry:- Regression: p/3
- Here,
mtry = 6is used
set.seed(1)
rf.boston <- randomForest(medv ~ ., data = Boston,
subset = train, mtry = 6, importance = TRUE)
yhat.rf <- predict(rf.boston, newdata = Boston[-train, ])
mean((yhat.rf - boston.test)^2)[1] 19.62021- Random forest test MSE ≈ 20.07
- Improves over bagging performance
- Use
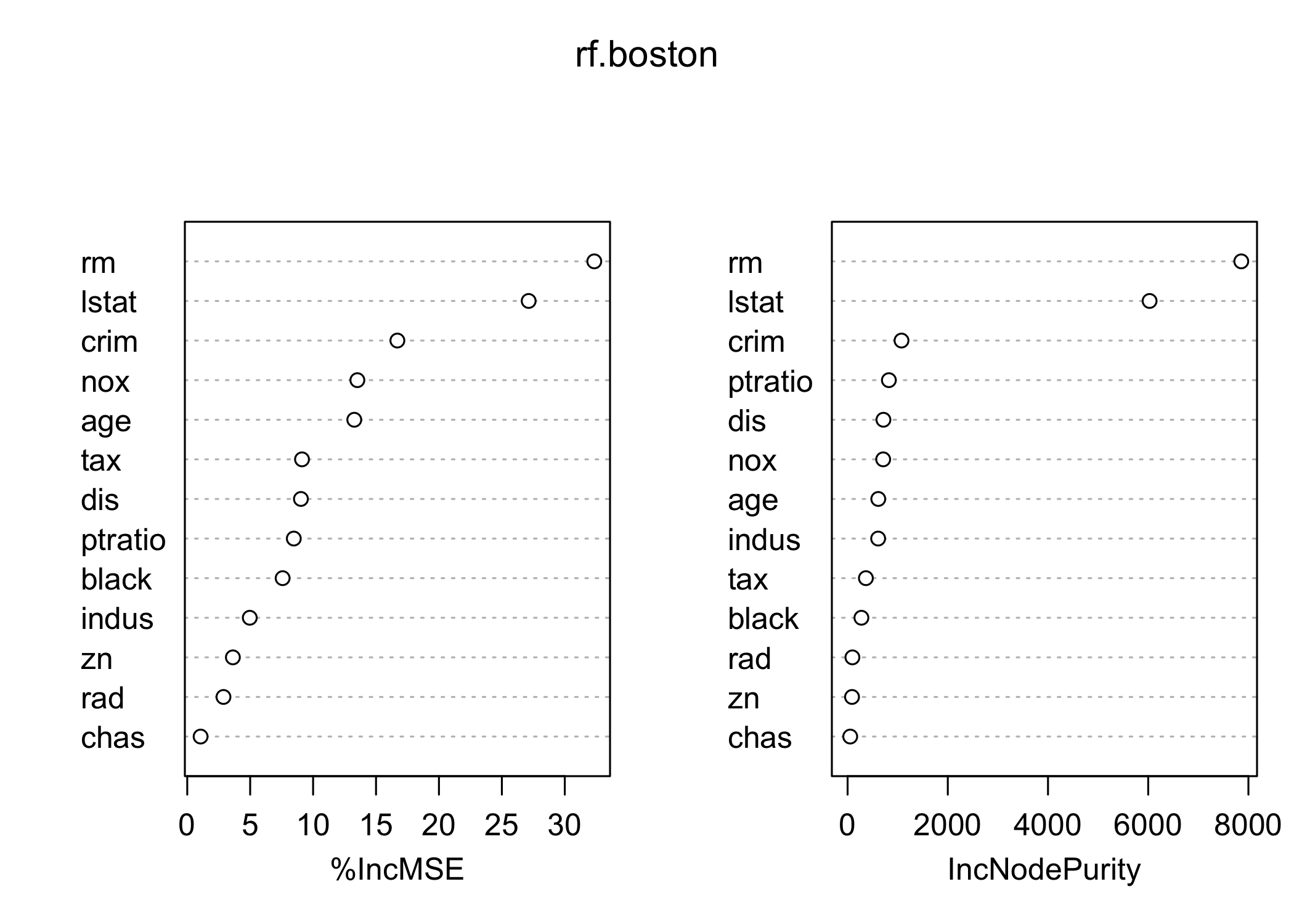
importance()to evaluate variable importance
%IncMSE IncNodePurity
crim 16.697017 1076.08786
zn 3.625784 88.35342
indus 4.968621 609.53356
chas 1.061432 52.21793
nox 13.518179 709.87339
rm 32.343305 7857.65451
age 13.272498 612.21424
dis 9.032477 714.94674
rad 2.878434 95.80598
tax 9.118801 364.92479
ptratio 8.467062 823.93341
black 7.579482 275.62272
lstat 27.129817 6027.63740- Two measures of variable importance:
- Mean decrease in accuracy (via permutation on out-of-bag samples)
- Mean decrease in node impurity (averaged over all trees)
- Most important variables in the random forest:
lstat(wealth / socioeconomic status)
rm(house size / number of rooms)
Boosting
- Use the
gbmpackage to fit boosted regression trees
- Apply
gbm()to theBostondataset
- Set
distribution = "gaussian"for regression problems
Loaded gbm 2.2.2This version of gbm is no longer under development. Consider transitioning to gbm3, https://github.com/gbm-developers/gbm3summary()provides:- Relative influence plot
- Numerical measures of variable importance
- Relative influence plot
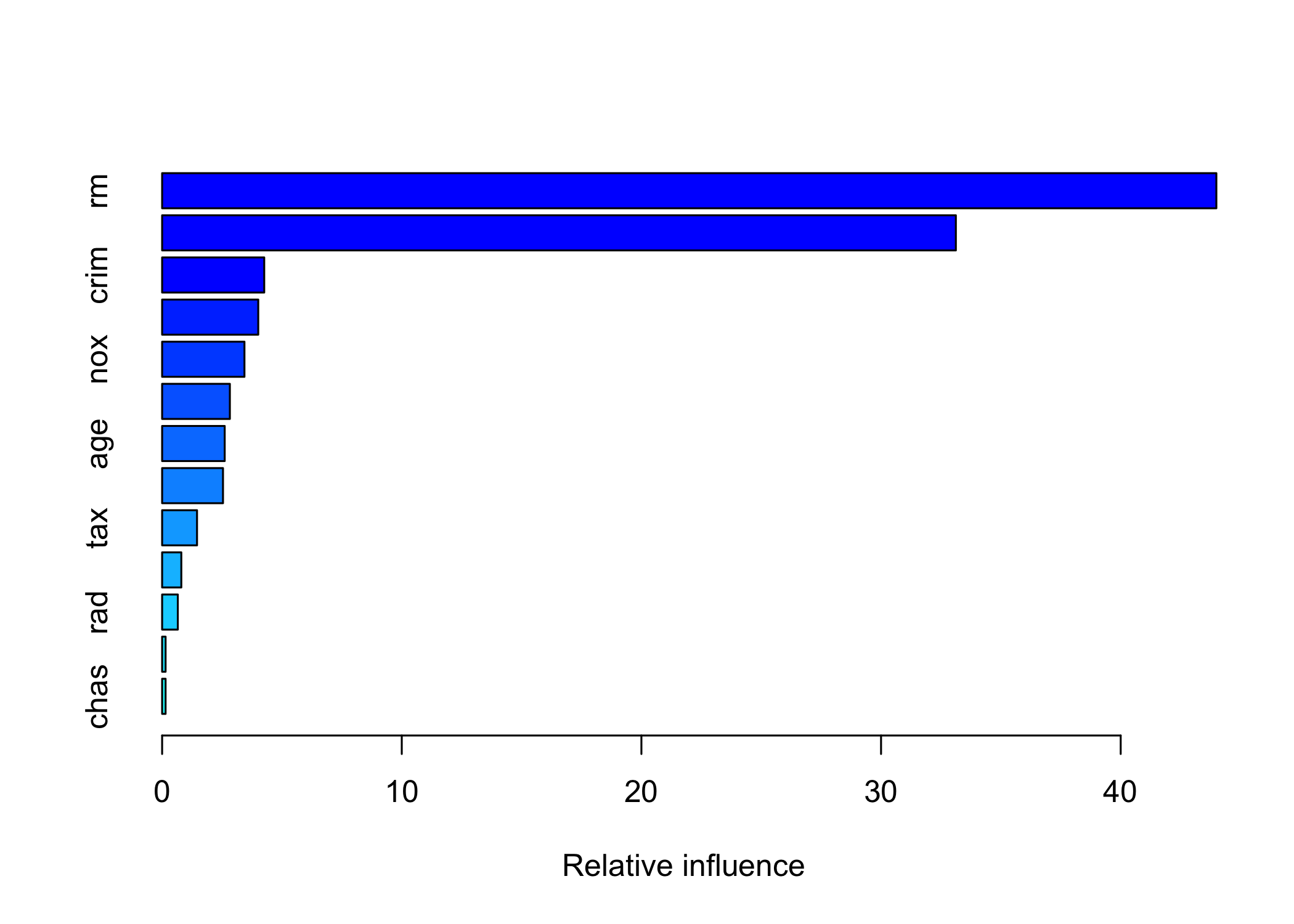
var rel.inf
rm rm 43.9919329
lstat lstat 33.1216941
crim crim 4.2604167
dis dis 4.0111090
nox nox 3.4353017
black black 2.8267554
age age 2.6113938
ptratio ptratio 2.5403035
tax tax 1.4565654
indus indus 0.8008740
rad rad 0.6546400
zn zn 0.1446149
chas chas 0.1443986lstatandrmare the most important variables
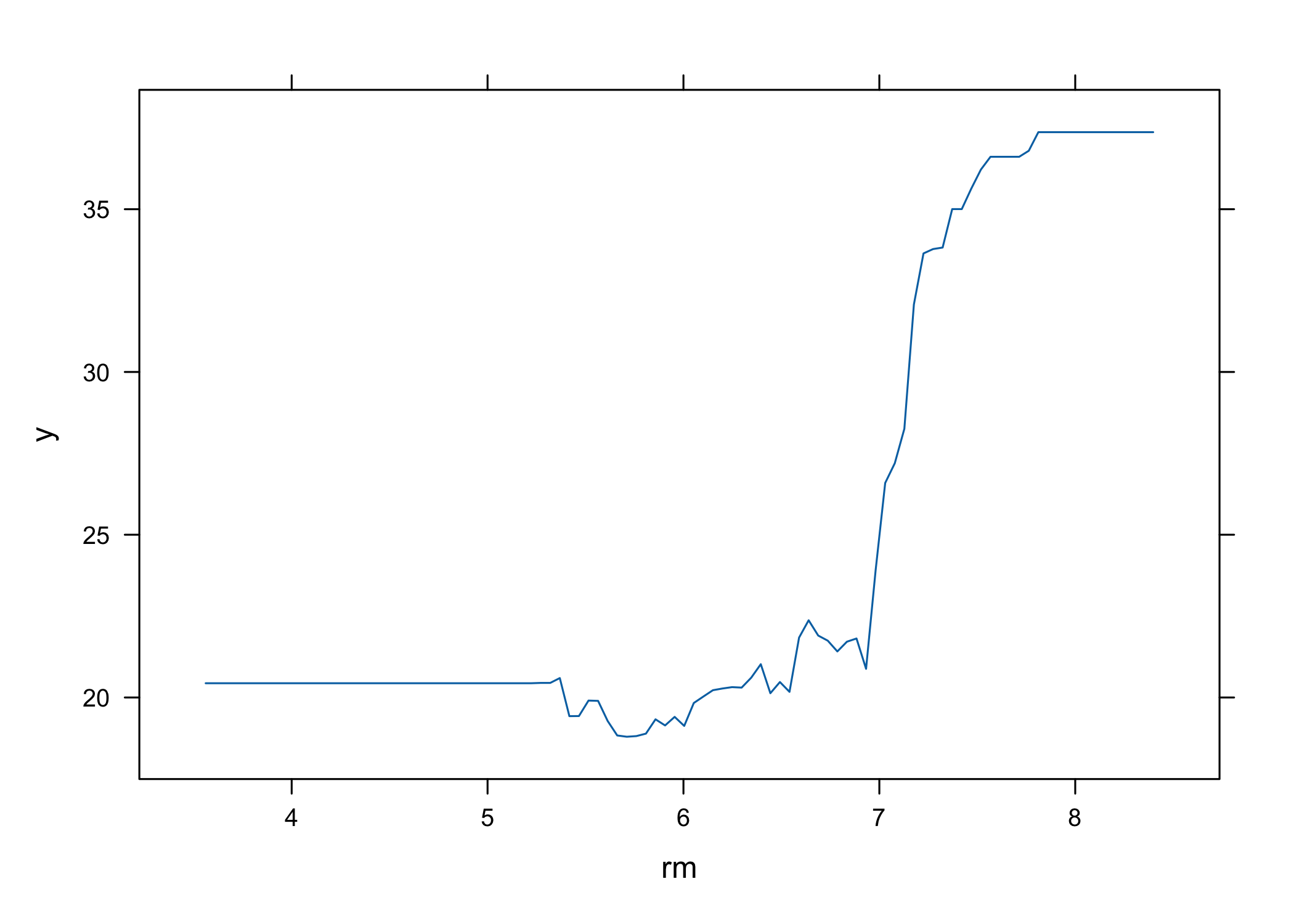
- Partial dependence plots can be generated for these variables
- Partial dependence plots show marginal effect of variables on response
- Effects observed:
- Higher
rm→ higher house prices
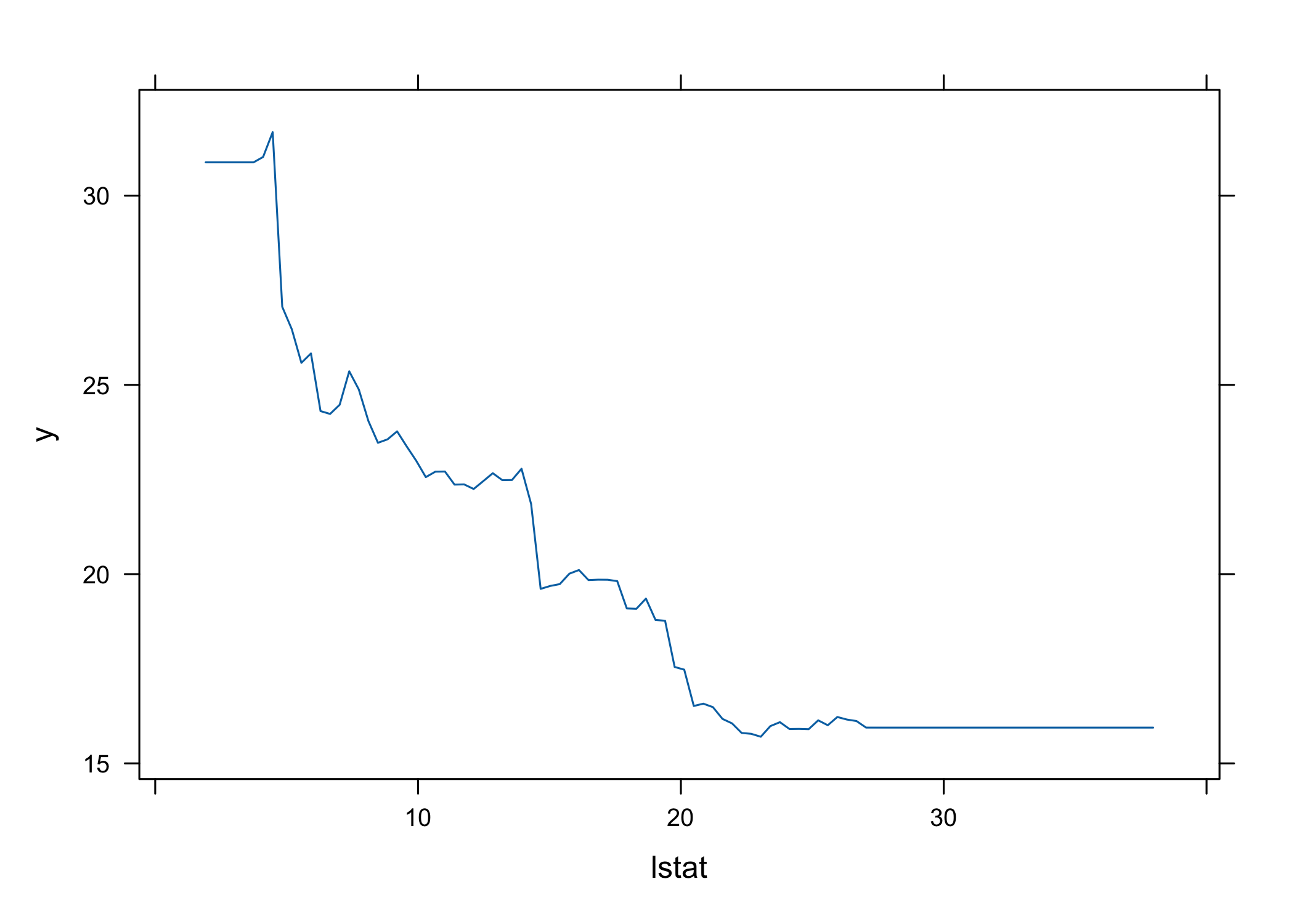
- Higher
lstat→ lower house prices
- Higher
- Use boosted model to predict
medvon the test set
yhat.boost <- predict(boost.boston,
newdata = Boston[-train, ], n.trees = 5000)
mean((yhat.boost - boston.test)^2)[1] 18.84709- Boosting test MSE ≈ 18.39
- Outperforms random forests and bagging
- Model can be tuned via shrinkage parameter \lambda
- Example: use \lambda = 0.2
boost.boston <- gbm(medv ~ ., data = Boston[train, ],
distribution = "gaussian", n.trees = 5000,
interaction.depth = 4, shrinkage = 0.2, verbose = FALSE)
yhat.boost <- predict(boost.boston,
newdata = Boston[-train, ], n.trees = 5000)
mean((yhat.boost - boston.test)^2)[1] 18.33455- Using \lambda = 0.2 results in lower test MSE than \lambda = 0.001